 |
Top : Molecular Biology : Protein : Protein Electrophoresis : SDS-PAGE : Protocol for SDS-PAGE Analysis
Protocol for SDS-PAGE Analysis | |
| Author: Arun Malik, Harnish, Priyanka Chopra, Stanzin Angm* | |
| Affiliation: National Agri-Food Biotechnology Institute, Mohali, Punjab, India | |
| Source: Protocol Online | |
| Date Added: Thu Feb 28 2013 | |
| Date Modified: Sun Mar 10 2013 | |
| Abstract: This protocol described the validated protocol in our lab for SDS-PAGE analysis of proteins. | |
PROTOCOL FOR SDS-PAGE ANALYSIS
Arun Malik, Harnish, Priyanka Chopra, Stanzin Angmo and Hariom Yadav[*]
National Agri-Food Biotechnology Institute, Mohali, Punjab, India
Introduction
SDS-PAGE technique broadly used in molecular biology, used to separate proteins according to their length of a linear polypeptide chain. The general electrophoresis technique cannot use to measure molecular weight of biological molecules but this technique mostly used to measure the molecular mass. The term electrophoresis means movement of charged particles to an electric field. In an electric field movement of proteins towards electrode of opposite charge. The separation of proteins according to their shape, size takes place.
SDS-PAGE conducted on casting the gels to avoid trouble and hazard of working with acryl amide. In this protocol presence of SDS and beta-mercaptoethanol which is used to denature the proteins in to their subunits.
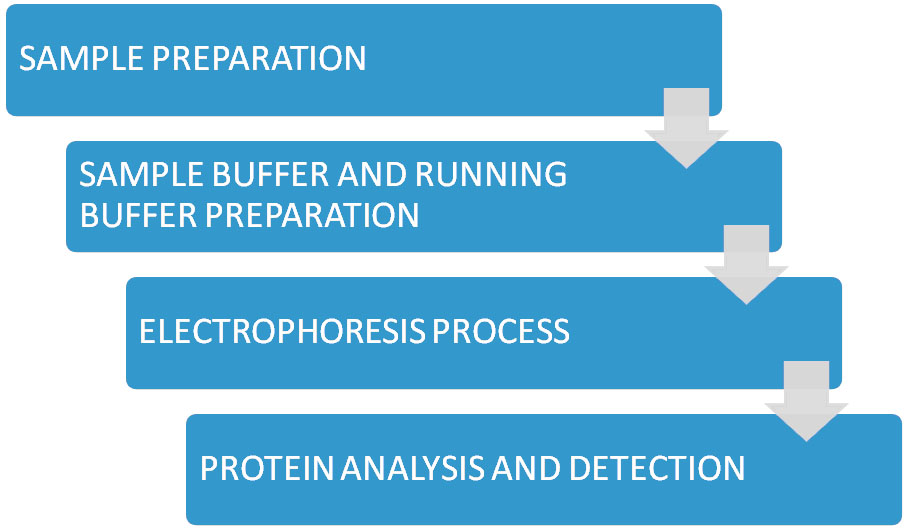
Figure. SDS-PAGE Process Flow
Reagents:
Stock solutions: resolving gel for 12% solution
- 30% Acrylamide/ Bis (37.5:1)
Take 29.2gm acrylamide + 0.8 gm N,N’- bis-methylene-acrylamide. Add to 100ml water to it.
- 1.5 M Tris-HCL, pH-6.8
Take 6gm Trizma Base. Adjust pH 8.8 with 6 N HCL. Add to 60ml water to it.
- 10% (w/v) SDS
To make 100 ml SDS, take 10gm SDS and 90 ml water add to it.
- Distilled water 3.35 ml.
- TEMED 5 microliters.
- 10% APS (Ammonium Persulphate) 50 microliters.
Materials
- PAGE Rigs includes spacers, combs, glass plates ( 10 X 20 cm).
- Protein sample.
- Sample buffer containing SDS and either sucrose or glycerol.
- 2-Mercaptoethanol.
- 1 X Running Buffer
- 5 X Sample Buffer
25 mM Tris-HCL
200 mM Glycine
0.1% (w/v) SDS
Approximately volume of water keeping below 1 liter that depends on electrophoresis system.
10% (w/v) SDS
10 mM Dithiothreitol or betamercapto-ethanol
20% (v/v) Glycerol
0.2 M Tris-HCL, Ph-6.8
0.05% (w/v) Bromophenolblue
Procedure
Gel pouring into miniprotein cassette
- Prepare resolving gel, and add all reagents to make gel except TEMED and ammonium persulfate (APS). Once TEMED and APS added then solution with get polymerize with in few minutes.
- Set the casting glass frames on casting strand. Pour the resolving gel between glass frames.
- After pouring gel, insert comb gently and before gel gets polymerize.
- Leave the polymerize until gel gets firm up to 25-30 minutes.
Electrophoresis
- Take the glass frames from casting strand and keep in to electrophoresis chamber.
- Fill the chamber with buffer and load the protein samples in to wells of gel then close the electrophoresis chamber.
- Connect the red and black electrical leads in to the matching colored electrodes of the electrode assembly.
- Put voltage around 120 volt and we can use 12% separating gel. Normally run the gel about 40-50 minutes till bromophenol dye has migrated to the bottom of gel.
- After migrating dye bottom of gel remove the electrode assembly.
Protein analysis and detection
Visualize the protein using Coomassie Brilliant Blue, silver stain, or we can use any other protein stain.
Notes
- Use TEMED and APS after mixing all reagents before few minutes of pouring the gel otherwise gel will gets solidify.
- Keeps glass frames exactly right upon each other otherwise gel will pour through the glass frames by keeping gasket on to right position on the cassette.
- For electrophoresis unit process always connects red and black leads to the matching colored electrodes of the electrode assembly.
[*] To whom correspondence should be made: yadavhariom@gmail.com