AZA treatment induces the genes but bisulfite sequencing does not reveal de-meth - (Jan/27/2014 )
I have treated my breast cancer cells with 5uM 5-AZA for 3 days and and checked out the expression of the genes previously known to be methylated in breast cancer. The expressions of these known genes were induced to around 10x after 5-AZA treatment.
After that I scanned a bunch of genes and found their expression to be induced to the levels ranging from 2x to more than 1000x. However, when I sequenced the promoter of a gene which was induced more than 1000 fold in AZA treated cells, I found out that the promoter region of the gene was not de-methylated at all (even in some cell lines, the methylation seems to be increased a little).
Another gene's promoter turned out to be in the same situation. How is this possible? Please share if you have any comments or suggestions.(My bisulfite sequncing primers do not include CpGs, and the bisulfite conversion seems to be allright according to QUMA software (>95%))
Thanks in advance.
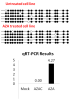
How did you do the normalization of your qPCR result? Maybe the reference gene is modulated by the treatment and this is the main cause of the observed difference? If you see 1000 fold change, you should also see this in your raw data already. Is the Ct value of your GOI significantly lower after treatment (with constant Ct for the reference gene)?
Thank you for your reply.
For the given example, the difference between AZA treated ct and the untreated ct is 11 ( Untreated ct is higher). I used GAPDH for the normalization and the Ct difference between AZA and untreated is 0.6 (AZA ct is higher). According to available microarray results, GAPDH does not change significantly with AZA treatment. Sİmilar case is present in other cell lines and other genes.
OK, so the effect seen is clearly caused by the upregulation of your GOI. That makes it a bit more difficult to find the answer to your problem.
Theoretically, all kinds of mechanisms are probable: the treatment may result in a different state of the chromatin in the region where your GOI is, making it more accessible to all kinds of transcription factors. Alternatively, the treatment may upregulate a transcription factor that activates your GOI, regardless of the methylation status. How sure are you that the DNA fragments you checked for methylation are the real and only promoters? You state that the genes were previously shown to be methylated in breast cancer. Did these publications also use the same cell lines as you did or were these different? If available, I would first try to mimick the published experiments to validate your method.
Good answers so far. My only $0.02 is that I have heard bisulfite sequencing can be rather tricky. I have done it myself, so only have secondhand info. You may want to try verifying the accuracy of this part of the experiment.
Thank you for your comments, we also consider that this may be a secondary effect of AZA. Do you know any other global effect of AZA which is NOT due to DNA de-methylation?
The problem is that, we expect AZA to cause demethylation throughout the genome (may be not completely but still at least most of it), but the promoter regions of my GOI and others we sequenced are still highly methylated in all AZA treated cell lines. And we know that AZA gets incorporated ito the genome during DNA replication, so I assume 3D structures of the chromosomes should not interfere where specifically AZA acts or not.Thus, we think it is not just by chance that these regions that we sequenced remained methylated, but that the de-methylation of the genome may not be successful (though we used the concentration and duration according to reference articles. And I selected the promoter region according to the previous articles studying my GOI).
So, if my GOIs were induced by some transcription factors as you suppose but not by de-methylation, what do you think caused the induction of this (or these) transcription factor(s) other than de-methylation at the beginning? Or do you assume that de-methylation occured in the promoter region(s) of this (or these) specific transcription factor(s) only? OR, do you think something is wrong with my point of view?
For the bisulfite sequencing part, we sequenced lots of promoter regions, and the promoter region was unmethylated in some cell lines or tissues while methylated in others. So we do not think that there is a bias caused by bisulfite sequencing method. Can you suggest anything else to verify the bisulfite sequencing experiment?
Thank you very much again four your help.
I'm not aware of other global effects of AZA than DNA demethylation. Since you don't see any demethylation, are you sure the AZA is what you think it is? Is it freshly ordered or an old stock? How do you dissolve it? It's been quite some years since I worked with that, but I seem to remember that it was quite unstable once dissolved and that I stored it at -80 °C. Do you add one 'shot' of AZA in the beginning after seeding the cells or do you add it also after some time (I seem to remember that in the culture medium it was also quite unstable).
The untreated control: do you seed these out at the same time as you treated cells? Does the AZA have growth retarding properties in your cell line and maybe the control is confluent after a couple of days and the treated are not? Differences in confluency can also have a big difference on expression of some genes. Probably you already thought of all these things, but I'm trying to find a technical explanation for the apparent discrepancy, cos that's easier to solve. For a biological explanation, it's more in the realm of guessing and difficult to investigate. The TF possibility I mentioned would indeed be as you said, it's promoter demethylated and as a result, upregulation of your GOI, but that's just speculating ...
Thanks a lot again.
AZA was freshly dissolved in DMSO (to be 22mM), aliquoted at one-use amounts and were kept at -20, and was used in a few weeks. It is 5-aza-2'deoxycytidine from Sigma. We changed the medium everyday, and added fresh AZA at each medium change. We performed cytotoxicity test before the experiment and chose the concentration which did not retard the growth of the cells significantly.
The DMSO treated control cells were seeded and collected at the exactly same times with AZA treated cells, and the DMSO added in these cells were exactly in the same amount as in AZA treated cells (and the medium was refreshed everyday as well).
As I had assumed, you already controlled for all the points I was thinking about so I'm sorry but I don't have any more suggestions to make ...
In our recent paper, we found that hypermethylated promoter of our gene was not demethylated by 5uM DAC (and even more stringent conditions). It's been already published, that DAC doesn't demethylate all hypermethylated regions, but the reason for this is not exactly known.
So this may explain, why your genes are not demethylated, if you don't have some error in PCR or bis-Seq (though this should be more of an exemption).
The reason why are upregulated anyway may be, it's not related to their methylation status.
Actually, we found, that not methylation but acetylation status of histones is more predictable of the expression regulation (by treating cells with DAC and trichostatin). Though methylation should be generaly predictor of condensed chromatin, it is just a mark, and probably regions can be "marked" with methylation but can be actively transcribing inspite of that.
You can do CHIP for histone activating and represing marks on your genes to test this.
