Electrophoresis after PCR : too many bands - (May/12/2014 )
Hello,
I'm new on the forum, and my english isn't so good, sorry for that !
We have problems understanding our gel electrophoresis after PCR. We have too many bands.
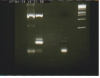
On the right side of figure we see the ladder. We try to understand the bands on the two first columns. We think that the two most intense bands corresponds to our PCR-Products.
What could be the others bands ? The plasmid ? The primers? Degradation of DNA?
We used PCR with phusion HF buffer, phusion DNA polymerase, forward and reverse primer, dNTPs, plasmid and DMSO. The sequence we want to amplify contains code for YFP and Cerulean proteins.
Thank you in advance !
Have nice day
Johanna
What band size do you expect? Most intense bands not necessarily means band of your interest.
27 kDa for YFP and 26.7 for Cerulean
I want to add that for the first column we should obtain product of PCR YFP and for the second column Cerulean.
Thank you for helping me.
As you are running PCR product it should be expressed in base pairs(bp). Proteins are generally denoted by kDa.
Tell exact size of your PCR product in bp. If you find some difficulty in directly searching it use https://www.genscript.com/conversion.html
What ladder do you used? - 50bp, 100bp or 1kb?
Ok, thank you for these informations.
So the size of the YFP is 717 bp and for Cerulean 714 bp. The ladder is 1kb.
Thank you
Bands in bottom of lanes are primer dimers, it can be found below 50 bp regions. If you compare ladder bands with your protein bands, you are right bright bands are bands of your interest. YFP lane upper band could be plasmid band, but at higher plasmid concentration you rarely get PCR amplification so other possibility is that it is nonspecififc amplification. Bands at random location other than band of your interest are nonspecific amplificatiion. To get specific bands, you can adjust various conditions as temperature, Salt concentration, template concentration etc.
Thank you so much !
Have a nice day,
Johanna
