Diffuse rRNA bands on RNA denaturing (formaldehyde) gel - (Aug/28/2013 )
Hi lab people!
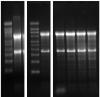
I am having a problem with my agarose denaturing RNA gels. I started using the standard protocol with 2,2M of formaldehyde in the gel, but got suspicious it was too concentrated and tried to go as low as 1M without any improvements. Its suppose to be used for northern blotting so I would like to have sharper bands. It is particularly funny since in my previous lab it use to work just fine. Following is the protocol summary and gel picture (loaded 5ug of total RNA).
Anyone has any suggestions or solved a similar problem before? Thanks a lot in advance!
Protocol:
Gel: 1g agarose + 80 mL water. Heat-dissolve, adjust volume to compensate evaporated water, cool to around 60°C, add 10mL 10X MOPS and 10mL of formaldehyde 37% (12,3M).
Denaturing loading buffer (1mL always fresh): 500uL formamide, 170uL formaldehyde (same as above), 130ul 10X MOPS, 200uL LB RNA.
LB RNA: 30% Ficoll, 1mM EDTA, 100ug/mL Ethidium Bromide, Bromophenol Blue.
For loading the gel I use 1 volume of RNA mixed with 3 volumes of denaturing loading buffer, heat at 65°C for 5 minutes, snap cool in water-ice and load on 10 min. pre-runned gel.
Result attached!
Thanks again for any suggestion!!!
are you using the same brand and grade of agarose as in your previous lab? what about the other components?
Not at all. Most components are different from the ones I used previously, but on the other hand, the ones I'm using now are supposedly of higher grade (mostly all Sigma, mol biol grade, RNase-free stuff). I really don't have a clear explanation for this. I was kind of expecting someone to say: "oh, it happened before to me, you're using old formaldehyde or bad formamide". Today I'll do another test using all reagents from another lab. Lets see how it comes out...
Thanks!
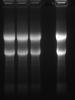
So... Did a test with everything new, and tested 6 alterations on the denaturing loading buffer. All look kind of similar, but lane two is the variation with no formaldehyde at all (just formamide, MOPS and LB). Looks like its the formaldehyde that's messing up the sharpness of my RNA bands, however, I don't have an explanation for the faster migration of this sample. RNA not completely denatured perhaps? In addition, I believe that if I take the formaldehyde off the gel recipe, then Ill get full renaturation of the RNA during the run and a bad Northern blotting... Maybe I should do the Northern as is or switch to glyoxal gels.
you can omit the formaldehyde from the loading buffer and leave it in the gel (if it improves appearance).
it's possible that the formaldehyde is not in good shape, it tends to polymerize. you may want to try a fresh lot of formaldehyde.
formamide also decomposes readily. you may be seeing problems with it. you can try fresh or try to omit it from the loading buffer.
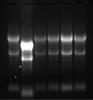
Solved the mystery a while ago and felt I should post it for other people not to follow the same route: Not formaldehyde, not formamide, not my RNAs, in fact, agarose was the main issue. Looks like you need to use low EEO agarose for RNA gels. It’s some chemical properties that basically influence how much the agarose polymer interacts with your migrating nucleic acids. Here is Sigmas explanation: "Electroendosmosis (EEO) - a movement of liquid through the gel. Anionic groups in an agarose gel are affixed to the matrix and cannot move, but dissociable counter cations can migrate toward the cathode in the matrix, giving rise to EEO. Since electrophoretic movement of biopolymers is usually toward the anode, EEO can disrupt separations because of internal convection."
Now, cheer up to "all purpose" UltraPure™ Agarose from Life/Invitrogen. It gives you crappiest, unresolved denaturing RNA gels. At least in my hands... If they are willing to send me a batch that works, I'll gladly change my opinion and post the results.
Below are the gel pics. Left is using UltraPure™ Agarose, then on the right some I did with a borrowed generic brand (cheaper) but low EEO. Ladder is 4ug Millenium RNA ladder.
Cheers!