Recombinant Plasmid is not being amplified in transformed cells - (Aug/05/2013 )
Hi Dear All,
For the past few weeks, I was trying to insert Hygromycin resistance cassette (eukaryotic expression) into a lentiviral vector.
I successfully inserted the sequence (~1.4kb) into a 7.7kb vector both by overlap extension PCR and by restriction enzyme cloning as I confirmed ligation/Overlap extension PCR by amplifying insert from recombinant E.coli DH5a colonies by colony PCR.
I verified recombination by amplifying insert as well as plasmid-insert junction sequence by colony PCR
Everything goes well until colony PCR but after that I lose recombinant plasmid and only get recipient vector from minipreps.
Why do I lose recombinant plasmid upon culturing DH5a cells? Is the hygromycin resistance cassette (HSV TK promoter-HygromycinR-HSV TK poly A signal) toxic to DH5a cells?
I inserted Hygromycin resistant cassette at different positions by both methods without interfering with Origin of replication or ampicillinR region.
Desperately looking forward to hear any positive feedback.
Thanks in advance.
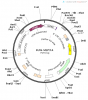
Recipient vector and recombinant vectors images are attached herewith this post.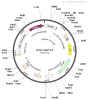




As you are working from colony PCR it is possible that you are amplifying the ligation mix, not necessarily the plasmid with insert. How are you doing the colony PCR? Are you amplifying using genes anchored in the vector or one in the vector and one in the insert or insert only?
It is possible that the insert is toxic, but it is unlikely as people have done these sorts of inserts before.
In my first colony PCR, forward and reverse primers were specific for vector and insert, respectively. So they amplified only recombinant product. As is shown in the attached colony PCR image, not all colonies are positive and there is no non-specific amplification either.
In overlap extension PCR cloned plasmids, I only used insert specific primers so we can deduce amplification of transformed insert.
What should I do to tackle this issue?
