HIV gene- PCR - (Dec/06/2012 )
Hi all,
I have been trying to amplify the HIV gene from HIV infected patients' genomic DNA. I am not getting the respective band, instead I'm getting a non-specific band with all the samples.![]() I checked the template DNA with other genes, its working fine. I checked the primers by using them with PNL4-3 as template, I got the exact band length. I tried changing the cycling conditions, template quantity, MgCl2 conc., tried adding DMSO (2%), everything possible, still no use.. What else can I try doing so that I get the required amplification..??
I checked the template DNA with other genes, its working fine. I checked the primers by using them with PNL4-3 as template, I got the exact band length. I tried changing the cycling conditions, template quantity, MgCl2 conc., tried adding DMSO (2%), everything possible, still no use.. What else can I try doing so that I get the required amplification..??![]()
Suggestions would be greatly appreciated...
Thank you in advance.. ![]()
![]()
Hi Paridhy,
Welcome to BioForum.
It would be great if you could post some information.
1. Gel picture of your unspecific bands that you are getting.
2. How are you extracting gDNA from patient samples ( Often blood proteins (contaminants) do not let your PCR work. Hence the non specific amplification. Am guessing here that your positive control template is purified with a protocol different from the one you are using for extracting gDNA.
Hi Ameya,
The control I used was a vector that was readily available. I used QIAmp DNA isolation kit (mini) to isolate the gDNA. So, I think, the quality of the DNA would be fine.![]() I have also attached the gel pic (1st well: 100bp ladder; 2nd well: The PCR product, shows the nonspecific product; 3rd well: The positive control, HIVpNL4-3; 4th well: The negative control).
I have also attached the gel pic (1st well: 100bp ladder; 2nd well: The PCR product, shows the nonspecific product; 3rd well: The positive control, HIVpNL4-3; 4th well: The negative control).
Thanks.. ![]()
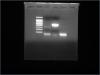
If you used plasmid as a control (and next time I would probably use much less of it, 100 or even 1000x dilution, plasmid DNA has high copy number) it's not a complex template so it won't show nonspecific binding to other targets in human genome. At this picture the primers show very high dimer formation so I wouldn'd consider them very good. If everything fails you may try to redesign them.
Do you have a negative human sample to compare? It's also possible that the "nonspecific" band is actually specific and only different length than plasmid control. (I assume negative control is just water).
