questions about shearing chromatin analysis - (Apr/19/2012 )
Hi all,
I did the chromatin shearing with Biruptor.( condition is Power "high". 10 times for 30 seconds at 30 second intervals.) The gel pic looks like a lit bit strange. I could not find the genomic DNA band on unsheared samples. Who could you give me some idea.
In the gel pic, I have 5 different concentration cell samples. The sample marked U is that unsheared sample and marked S is that sheared samples. The unsheared samples i took them before put in Biruptor and directly load on the gel without add any RNAse and incubation. I need help to continue. If i have problem with cross-link or lysis buffer.
The lysis buffer recepie is
Lysis Buffer fresh reagent
Many thanks!
Ting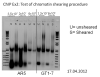
since you didnt reverse the crosslinked chromatin you can not expect your unsheard sample to run into the gel............my guess is that what you are primarily looking at is RNA in all of your lanes. You need to reverse the crosslinks and RNAse treat your samples to get any meaningful information on whether your shearing conditions are good or not..............your lysis buffer looks fine to me.
Hi chabraha,
I doubt my lysis step did not work and the shearing was ok.So I have repeat this lysis procedure again. This time i add the 20 gauge needle treatment and reverse cross-linking step. The result see the attachment. This time i could find the genomic DNA band in the unsheared lysis samples. So i think I could continue to next step, do you think so. Thanks:)
Sure.......I run my samples 6 times through a 20g needle and it definitley helps crack the nuclei and slightly shear the DNA....just be sure to RNAse your sample before you load it onto your gel...........otherwise you can't be sure whether you are looking at RNA or DNA.
