Ligation gel picture interpretation - (Feb/01/2012 )
Hello, I am trying to insert a 2 kb PCR fragment with EcoRI/XbaI sites into a pET32a vector (5.5 kb) that is digested with EcoRI/XbaI. I did a 3 hour ligation.
Control: Vector + ligase - - - - This gave approximately 50 colonies
1:3 Vector:Insert - - - - - - again around 50 colonies
1:8 Vector:Insert - - - - - around 60 colonies
I am worried about all of the self ligation that occurred and I doubt that any of my colonies on experimental plates are the clone I am looking for.
I ran some of the leftover ligation on a gel and would like some input on the multiple bands I observe in the vector + insert ligation reactions. I will be doing minipreps and XbaI/EcoRI digestions tomorrow so I will find out then if it worked for sure, but is the expected 7.5 kb vector+insert band in the smear?
lane 1: marker (12000, 8000, 6000, 5000, 4000...)
lane 2: control (vector + ligase)
lane 3: 1:3 vector:insert
lane 4: 1:8 vector:insert
Also, if you have any suggestions to limit self-ligation since my control plate had as many colonies as the actual ligation they would be appreciated. I have not used SAP or the likes for the vector since I have incompatible ends.
Thanks in advance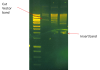
the digest doesn't seem to be complete. Try to digest it completely and run the digest longer on gel before purifying it for ligation.
I digested the vector overnight (~18 hours) and it seemed to give a very defined band on the gel that I excised. Are you saying since there are three bands in the vector + ligase (lane 2) the digest of the vector must have been incomplete? I was thinking the top band could possibly be a dimer of the vector
if digested completely, you should have one clear band for the vector alone control group.
UPDATE: It turned out I did have clones from this ligation reaction, but as scolix predicted the majority were uncut plasmid that was transformed. I screened 18 colonies and I had 3 positive clones.
