|
This is a cached page for the URL (http://www.fgsc.net/fgn/camb.html).
To see the most recent version of this page, please click here.
| | Protocol Online is not affiliated with the authors of this page nor responsible for its content.
About Cache | | |
An Ultra-fast method of DNA extraction from Neurospora An Ultra-fast method of DNA extraction from Neurospora
Edward B. Cambareri and John A. Kinsey - Department of Microbiology, Molecular Genetics and Immunology, University of Kansas Medical Center, Kansas City, KS 66160 We have found that the DNA extraction procedure of Metzenberg and Baitch (Neurospora Newsl. 28:20)/Stevens and Metzenberg (Neurospora Newsl. 29:27) while giving excellent yield and size of DNA, is somewhat cumbersome and also results in the occasional sample that proves to be uncuttable. We report here a simple method for growing Neurospora and for isolation of DNA that may be performed in two days from start to finish. The growth of mycelia in Petri plates (suggested by G. May) eliminates the need for large numbers of flasks when growing many cultures for DNA isolation.
1. Inoculate plastic Petri plates containing 25 ml of 1X Vogels, 2% sucrose, 0.04% Tween-80 (to prevent conidiation) (Zalokar, M. 1954. Arch. Biochem. Biophys. 50:71-80) plus any other required supplements. We use a loopful of conidia from a 5-10 day old tube. Incubate at 33°C. Handle the plates carefully to prevent spillage.
2. Harvest the mycelia by filtration after 36-48 h, rinse with distilled water and remove excess water by briefly pressing the pad between paper towels. Lyophilize for at least 2 h.
3. Grind the lyophilized pad in a 1.5 ml microfuge tube or a 15 ml conical tube using a metal rod or a spatula.
4. Add 1 ml of extraction buffer to the tube containing the mycelial powder and mix gently. Let stand at room temperature. for 20 min.
5. Transfer the homogenate to a microfuge tube and centrifuge in a microfuge for 5 min.
6. Recover supernatant and transfer to a tube containing 1 ml of diatomaceous earth (DE) in 6M Guanidine Thiocyanate. Let this stand at room temperature. for 2-3 min.
7. Decant into a spin column (either a 3 cc syringe body plugged with siliconized glass wool or a prefabricated spin column) and filter by aspiration. Rinse 3 times with 2 ml of 80% isopropanol. Continue aspiration for 2 min to dry out the DE. We use a standard 1 liter filter flask with a #8 stopper drilled with 5 small bore holes. Ten samples can be filtered simultaneously by using two flasks from one aspirator with a Y-junction. Commercial manifolds are also available.
8. Place each column into a 15 ml conical tube with the tip of the column resting inside a 0.5 ml microfuge tube at the bottom of the 15 ml tube. Add 150 ul of ddH20 at 65°C and spin in a tabletop centrifuge at maximum speed for 1 minute. Add an additional 150 µl and repeat.
Extraction Buffer
2M NaCl
0.4% Deoxycholic acid (sodium salt) Sigma D-6750
1.0% Brij 58 (polyoxyethylene 20 cetyl ether ) Sigma P-5884
DE/GTC
8.5 g of acid washed diatomaceous earth (Sigma D-5384), added to 100 ml of 6M Guanidine Thiocyanate (ultrapure, BMB)
Stable for approximately 6 months at room temperature.
The yield of DNA varies from 5-30 µg depending upon the amount of mycelium that is harvested. The quality of the DNA is high, with the size averaging greater than 20 kb. It digests to completion in 2 h, and has been used in Southern blots (Figure 1) and for PCR applications. The DNA is more dilute than in typical extraction procedures, but may easily be concentrated by lyophilizing or by precipitation.
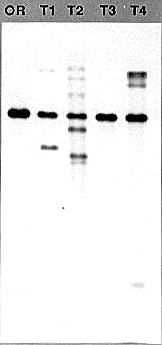
Figure 1 Genomic Southern blot analysis using DNA prepared with the Metzenberg procedure (OR, strain Oak Ridge 74-OR23-1VA) and DNAs prepared with the procedure described above (Transformants T1-T4). The visible bands range in size from>12 kb to 500 bp. One microgram of each of the DNAs was digested for 2h at 37°C with 5 units of HindIII in the manufacturers supplied buffer in a volume of 40 µl. The DNAs were fractionated overnight in a 0.7% agarose gel in 1X TAE buffer. After transfer to nylon membrane, the DNAs were probed with the transforming DNA fragment that was labeled by the random hexamer priming method. The probe is a 2.3 kb HindIII to BamHI fragment from linkage group VI, cloned from a strain where it forms a translocation junction with linkage group V just upstream of the am gene.
