In situ hybridisation to alpha satellite sequences (chromosome specific)
Notes
Alpha satellite sequences, whilst highly repetitive, are specific to each individual chromosome. These sequences flank the centromeres and can present a target measured in megabases. In this protocol a biotin or digoxigenin labelled DNA probe is detected using HRP-conjugated antibodies. The signal is visualised with diaminobenzidine (DAB). Normal, healthy nuclei show two spots. Aneusomic nuclei show 1, 3, 4 or more spots.
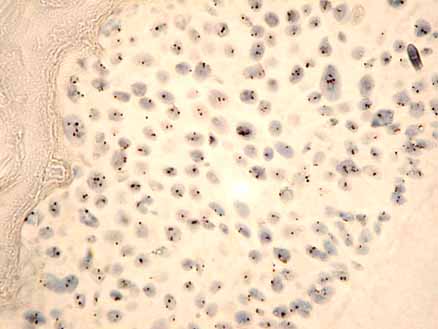
A 6mm section of a paraffin embedded raft culture hybridised to a chromosome 17 specific alpha satellite probe.
We buy our probes from Appligene/Oncor although they are very expensive and the telesales staff are rude.
Starting material
We have two types of starting material; (i) tissue culture cells attached to glass slides, (ii) sections taken from paraffin embedded tissue.
Tissue culture cells. We use multispot slides with 4 wells separated by PTFE coating. The slides are treated with APES prior to use (see below) . The cells are trypsinised and resuspended as usual then one or two drops are placed onto the end well of a labelled slide. The cells are allowed to attach overnight in a humid chamber in the CO2 incubator. Next day the slides are immersed in 10% neutral buffered formalin. After 2 days fixation the slides are rinsed in H2O and allowed to air dry. Store dry, dust-free.
Paraffin sections. Take 6mm sections onto APES treated slides (see below) and bake overnight at 60C. Dewax in Xylene, 2 x 5 minutes. Then immerse as follows; 100% ethanol 2 x 5 minutes, 90% Ethanol 5 minutes, 75% ethanol 5 minutes, H2O 2 x 5 minutes and allow to dry. Store dry, dust-free.
Typical Protocol
(1) Immerse the slides in 1M sodium thiocyanate at 80C for 10 minutes. Rinse in H2O and allow to dry.
This treatment strips away the histones.
If using cheap glass coplin jars, allow to heat up slowly with the water bath, otherwise they crack.
(2) Treat the slides with 4mg/ml pepsin at 37C for 30 minutes. Rinse in H2O, 2 x 5 minutes, and allow to dry.
This opens up the fixed (crosslinked) cytoplasm to allow probe access.
Make up 0.2M HCl [50mls H2O + 800ml conc. HCl], weigh out a little pepsin (Sigma P7012) into a universal and add the 0.2M HCl to make 4mg/ml. Place the slides flat in a humid chamber (eg a large square plastic petri dish) and drop the pepsin gently onto the slide. Then place in the 37C incubator.
This pepsin digestion is the most crucial step. Too little digestion and the probe will not be able to access the chromosomes. Too much and the morphology will all be chewed up.
(3) Prepare the hybridisation solution.
One well of a multispot slide needs 6ml of hyb with a 13mm circular coverslip.
A square 22mm coverslip uses 10-12ml.
For larger than this you can put two slides face to face (acting as each other's coverslip) with 30ml of hyb solution.
| Solution A | 6mls de-ionised formamide 2mls 20 x SSC 2mls H2O 2g dextran sulphate Store in a universal at room temperature | 25ml |
| De-ionised formamide | (Sigma F7508) | 15 ml |
| probe | eg. Appligene/Oncor #CP5040-B.5 (biotin labelled) or label your own probe (~10ng/ml) | 5 ml |
| Carrier DNA | Sheared salmon sperm DNA (10mg/ml) | 0.5 ml |
| H2O | _ | 2 ml |
| 20 x SSC | _ | 2.5 ml |
| TOTAL | 50 ml | |
(4) Apply an appropriate amount of hybridisation solution onto the slide and cover with a coverslip.
(5) Incubate at 80C for 10 minutes
This denatures the DNA
(6) Transfer to a humid chamber and incubate overnight at 37C
(7) Dip the slides into a coplin jar containing wash solution, allow the coverslips to slide off. Immerse the slides in formamide wash solution for 20 minutes at 42C.
We wash in formamide wash solution at 42C. The wash can be done without formamide at higher temperature (try 0.25 x SSC at 72C for 5 minutes)
Formamide wash solution: 60ml formamide TOXIC (Sigma F7503), 30ml H2O, 10ml 20 x SSC, 80 ml conc. HCl (to bring the pH to 7)
Do not allow the slides to dry at any stage until after DAB detection.
(8) Immerse briefly in PBS to wash, then incubate with blocking solution for 20 minutes at room temperature.
Lay the slides flat in a humid chamber and drop the solution onto them
Do not allow the slides to dry
Blocking solution: 100ml PBS, 2g BSA (Sigma A3059), 50 ml Tween 20.
I carefully dry around the tissue (or round the edge of the slide if I cannot see the tissue) with a paper towel to define a wet area for the various washes.
(9) Drain off the blocking solution and replace with mouse anti-biotin antibody diluted 1:100 in blocking solution for 40 minutes at 37C.
We currently use DAKO antibody #M0743, 100-200ml per slide.
(10) Wash with blocking solution for 10 minutes
(11) Incubate with rabbit anti-mouse HRP antibody diluted 1:80 for 40 minutes at 37C
We use DAKO P0260
(12) Wash with blocking solution for 10 minutes
(13) Incubate with swine anti-rabbit HRP antibody diluted 1:100 for 40 minutes at 37C
We use DAKO P0217
(14) Wash in PBS for 5 minutes
(15) Detect the signal using Diaminobenzidine (DAB)
I use DAB tablets (Sigma D4293) to make 5ml of DAB solution and then carefuuly drop the DAB onto the slides. DAB is carcinogenic so take care. The signal takes about 10 minutes to develop. If you're lucky you may be able to see tiny spots at low power under the microscope (take care not to dip the lens in the DAB solution)
Follow these URLs to learn about safe disposal of DAB;
. . . . . . . . http://www.safety.duke.edu/EnviroPrograms/Dab.htm
. . . . . . . . http://www.histology.to/DABfaq.html
(16) Rinse in tap water, stain the nuclei using haematoxylin, de-hydrate through the ethanol series and mount in DPX.
The length of the haematoxylin stain depends on the kind of haemaotoxylin and how much it has been used. Try 30 secs at first. Check under the low power microscope.
.
This is an alternative to poly-L-lysine treatment. The APES coating makes the slides more adhesive for fixed tissue and living cells.
Fill several big metal slide racks with slides (I do hundreds at a time), then treat each rack sequentially as follows;
(1) Immerse in 1% acid/alcohol 30 mins
Acid alcohol is [1% (v/v) concentrated HCl, 70% Ethanol, 29% H2O]
This is to clean the slides
(2) Rinse in water and allow to dry
First rinse in running tap water, then immerse in distilled water, then allow to dry.
(3) Immerse in acetone for 10 minutes
(4) Immerse in [2%(v/v) APES in acetone] for 5 minutes.
APES is aminopropyltriethoxysilane (Sigma A3648)
(5) Immerse briefly in 2 sequential rinses of distilled H2O
(6) Allow to dry and store in a dust-free container
Use within months not years