Determining the Direction of Replication Fork Movement
Bonita Brewer, October 28, 1991
For an in-depth review of the method, see Friedman, K. and B. Brewer (1995) Analysis of replication intermediates by two-dimensional agarose gel electrophoresis. Meth. Enzymol. 262: 613-627.
Two dimensional agarose gels have proven to be very useful for identifying restriction fragments that contain replication origins and termini. However, the majority of genomic restriction fragments are replicated by a single fork passing through them. These fragments can be informative if the direction of fork movement can be determined. A simple modification of our 2-D gel procedure, that uses non-denaturing conditions for both dimensions (Brewer and Fangman, 1987, Cell 51:463), permits determination of the direction of fork movement without interference from nicked or broken DNA.
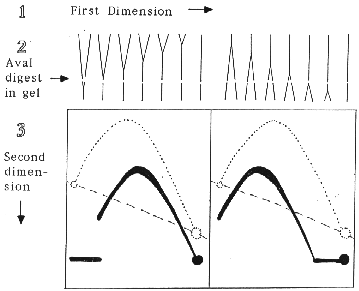
As an example of the procedure, consider an XbaI restriction fragment that is replicated by a single replication fork and has an AvaI site near one end. The XbaI cleaved replication intermediates are separated by mass in the first dimension of electrophoresis. The gel lane is excised, the DNA digested in the gel with AvaI and subjected to the second dimension electrophoresis where separation is by both mass and structure. A Southern blot probed for the larger XbaI-AvaI fragment will show that the detected replication intermediates, while reduced in size, still generate an arc of simple-Ys. However, the position of the 2-D pattern with respect to the arc of full-length XbaI fragments depends upon whether AvaI cleavage removes the first or the last end of the fragment to be replicated. The expected results are illustrated below.
Procedure for restriction enzyme cleavage in gelo. Adapted from the procedure published in the New England Biolabs Catalog (1988-1989; p. 142). The volumes and times recommended here are not necessarily optimal, since I haven't tried reducing them. Note that some restriction enzymes are inhibited by the presence of contaminants in agarose. As a trial run, I recommend cutting lambda DNA, for example, with the enzyme of choice in various brands of agarose. The enzymes that cut to completion in Beckman LE agarose are AvaI, Bam HI, BglII, EcoRV, NcoI, PstI, PvuII, SacII, SnaBI, SpeI, StuI, XbaI and XhoI. (There are likely to be many others that also cut to completion--see The NEB Transcript, Vol. 3 No. 1, March, 1991, pg 8-9.)
1. After staining and photographing the first dimension gel, excise the lane and place it in a disposable pipette reagent reservoir from Costar (#4870; Cambridge MA). These plastic boats have a wedge shaped reservoir that is 12 cm long. Fill up the container with 10 mM Tris (pH 8.0), 0.1 mM EDTA. Incubate at room temperature for 30 minutes with gentle agitation. Repeat this step once.
2. Make up 250 ml of restriction enzyme buffer according to the manufacturer's recipe. Be sure to include BSA but do not include spermidine in the buffer as it might alter the mobility of the DNA in the second dimension. Drain the Tris-EDTA from the gel piece and fill the container with the restriction buffer. Incubate 1 hr at room temperature, with gentle agitation. Repeat this step once.
3. Drain the restriction buffer from the gel slice. Remove excess buffer with a pasteur pipette and blot the gel slab with a kimwipe. Pipette directly onto the surface of the gel, 10 to 20 microliters of enzyme. Seal the reservoir with parafilm and incubate at 37o. (Alternatively, set the reservoir in a glass blaking dish with water in the bottom and cover with saran wrap.) Digest 4 hours (or more).
4. Fill the reservoir with 10 mM Tris, 1 mM EDTA. Agitate gently at room temperature 30 minutes.
5. The gel is now ready for the second dimension of electrophoresis. Beware that the DNA fragments are now smaller and thus the second dimension may need to be a higher concentration of agarose and/or may need to be run for a shorter time. The ethidium bromide stained profile may still contain a prominant arc of linears in the smaller size range; the larger fragments are more likely to have a site for the second enzyme and thus they will be distributed in a haze below the arc of linears. When analyzing the yeast genome it is possible to see single copy sequences as individual spots in this portion of the gel. In addition, the repeated sequences (rDNA and 2 micron) can provide a visual test for the apparent completeness of the second digest.
Return to top of page
Questions? Contact us